Amine coupling
Amine coupling makes use of the N-terminus and ε-amino groups of lysine residues of the ligand. Below a typical sensorgram of the immobilization of a ligand to a CM5 sensor chip using the amine coupling procedure. The numbered points refer to the different stages in the immobilization procedure.
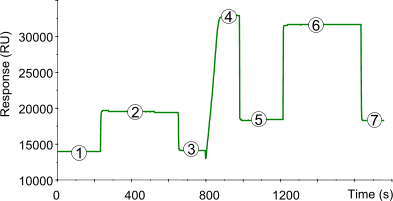
| 1) | Baseline for the unmodified sensor chip surface with continuous flow (5 µl/min). |
| 2) | 35 µl injection of NHS/EDC to activate the surface by modification of the carboxymethyl groups to N-Hydroxysuccinimide esters. |
| 3) | Baseline after activation. Activation of the surface has only a very slight effect on the SPR signal (100–200 RU). |
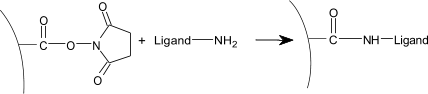
| 4) | Injection of ligand (10–200 µg/ml) leads to electrostatic attraction and coupling to the surface matrix. At this point, the ligand solution is still in contact with the sensor surface, and response includes both immobilized and non-covalently bound ligand. The N-Hydroxysuccinimide esters react spontaneously with the amines on the ligand to form covalent links (1). |
| 5) | Immobilized ligand before deactivation. The ligand has passed the sensor surface and most of the protein that is not covalently bound is eluted. |
| 6) | Deactivation of unreacted NHS-esters using 35 µl 1 M ethanolamine hydrochlorid adjusted to pH 8.5 with NaOH. The increased SPR signal is due to a change in the bulk refractive index. The deactivation process also removes any remaining electrostatically bound ligand. |
| 7) | Point 7 minus Point 3 gives the amount of immobilized ligand after deactivation. |
Amine coupling is the first choice with new molecules to couple. However acidic ligands (pI < 3.5) are difficult to immobilize. Also when the free amine groups are in the biological active site, one of the other chemistries must be investigated.
References
| (1) | Johnsson, B. et al Immobilization of proteins to a carboxymethyldextran-modified gold surface for biospecific interaction analysis in surface plasmon resonance sensors. Analytical Biochemistry 198: 268-277; (1991). |